 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 866329 | ||||||||
| Common Name | ARALYDRAFT_665773 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 307aa MW: 35131.6 Da PI: 7.078 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 48.1 | 2.7e-15 | 14 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT.-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg.tWktIartmgkgRtlkqcksrwqkyl 48
+g+W++eEd +l+ ++++ G+g +W + ++++g++R++k+c++rw++yl
866329 14 KGPWSPEEDAKLKSYIEKSGTGgNWIALPQKIGLKRCGKSCRLRWLNYL 62
79*********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 36.8 | 9.2e-12 | 69 | 111 | 2 | 46 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
g +++eE+ ++ + G++ W+ Ia+ ++ gRt++++k++w++
866329 69 GGFSEEEENIICSLYLTIGSR-WSIIAAQLP-GRTDNDIKNYWNT 111
569******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.987 | 9 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 4.49E-26 | 11 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.0E-12 | 13 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-14 | 14 | 62 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 6.1E-24 | 15 | 69 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.16E-8 | 16 | 62 | No hit | No description |
| PROSITE profile | PS51294 | 20.442 | 63 | 117 | IPR017930 | Myb domain |
| SMART | SM00717 | 1.5E-10 | 67 | 115 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-10 | 69 | 111 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-23 | 70 | 118 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 5.33E-7 | 71 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 307 aa Download sequence Send to blast |
MGRAPCCDKA NVKKGPWSPE EDAKLKSYIE KSGTGGNWIA LPQKIGLKRC GKSCRLRWLN 60 YLRPNIKHGG FSEEEENIIC SLYLTIGSRW SIIAAQLPGR TDNDIKNYWN TRLKKKLINK 120 QRKELQEACM EQQEMMVMMK RQQQQQQIQT SFMMRQDQTM FTWPLQHHHD QVPTLFMDKT 180 NSFCDQEDAK PVIKIEDQEL ERTNPHHHQD SMTNAFDHLS FSQLLLDPNH NHLGSGEGFS 240 MTSILSANAN PPLLNTSIND HQWFGNFQAE TINLFSGAST STSADQSTIS WEDISSLVYS 300 DSKQFC* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 8e-24 | 14 | 117 | 7 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator (By similarity). Positively regulates axillary meristems (AMs) formation and development, especially during inflorescence. {ECO:0000250, ECO:0000269|PubMed:16461581}. | |||||
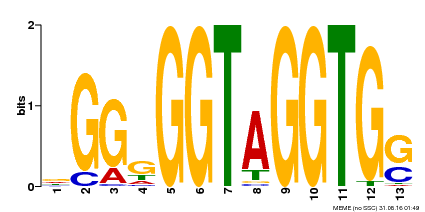
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00089 | SELEX | Transfer from AT3G49690 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 866329 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY089148 | 0.0 | AY089148.1 Arabidopsis thaliana clone 38611 mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002875991.1 | 0.0 | transcription factor RAX3 | ||||
| Swissprot | Q9M2Y9 | 0.0 | RAX3_ARATH; Transcription factor RAX3 | ||||
| TrEMBL | D7LSM9 | 0.0 | D7LSM9_ARALL; Predicted protein | ||||
| STRING | Al_scaffold_0005_1808 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2941 | 25 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G49690.1 | 0.0 | myb domain protein 84 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 866329 |
| Entrez Gene | 9312059 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



