 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 891746 | ||||||||
| Common Name | ARALYDRAFT_891746 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 311aa MW: 34188.4 Da PI: 9.3226 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 23.5 | 1.3e-07 | 7 | 54 | 3 | 45 |
SS-HHHHHHHHHHHHHTTTT......-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg......tWktIartmgkgRtlkqcksrwq 45
W+ eE++ + +a++++ + +W++ a+ ++ + l+++k ++q
891746 7 TWSREEEKAFENAIALHCVEeeitedQWNKMASLVP-SKALEEVKKHYQ 54
5****************999****************.***********9 PP
| |||||||
| 2 | Myb_DNA-binding | 38.8 | 2.1e-12 | 132 | 176 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r + Rt+ q+ s+ qky
891746 132 PWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPTQVASHAQKY 176
7*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 7.175 | 3 | 61 | IPR017884 | SANT domain |
| SMART | SM00717 | 0.0018 | 4 | 59 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.09E-6 | 7 | 57 | No hit | No description |
| SuperFamily | SSF46689 | 1.94E-8 | 8 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.873 | 125 | 181 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.9E-17 | 127 | 182 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.3E-10 | 129 | 179 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 4.4E-17 | 129 | 179 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.7E-11 | 130 | 175 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 7.7E-11 | 132 | 176 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.10E-9 | 132 | 177 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 311 aa Download sequence Send to blast |
MESVVATWSR EEEKAFENAI ALHCVEEEIT EDQWNKMASL VPSKALEEVK KHYQILLEDV 60 KAIENGQVPL PRYHHRKGLI VDEAATSPAN RDSHSSGSSE KKPNPGTSGI SSSNGGRSGG 120 SRAEQERRKG IPWTEEEHRL FLLGLDKFGK GDWRSISRNF VISRTPTQVA SHAQKYFIRL 180 NSMNRDRRRS SIHDITTVNN QAPTVTGGQQ QQQVVKHRPA QPQPQPQPQP QPHHPPTMAG 240 LGMYGGAPVG QPIIAPPDHM GSAVGTPVML PPPMGTHHHH HHHLGVAPYA VPAYPVPPLP 300 QQHPAPSTMH * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 3e-13 | 8 | 78 | 11 | 78 | RADIALIS |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
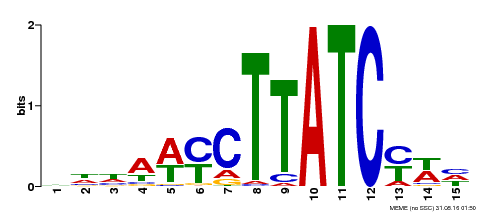
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 891746 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY086906 | 0.0 | AY086906.1 Arabidopsis thaliana clone 29302 mRNA, complete sequence. | |||
| GenBank | AY519528 | 0.0 | AY519528.1 Arabidopsis thaliana MYB transcription factor (At1g49010) mRNA, complete cds. | |||
| GenBank | BT024862 | 0.0 | BT024862.1 Arabidopsis thaliana At1g49010 gene, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002891472.1 | 0.0 | transcription factor MYBS1 | ||||
| TrEMBL | D7KDZ5 | 0.0 | D7KDZ5_ARALL; Uncharacterized protein | ||||
| STRING | scaffold_104540.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM10304 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49010.1 | 1e-150 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 891746 |
| Entrez Gene | 9330215 |



