 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | BGIOSGA017051-PA | ||||||||
| Common Name | OsI_17252 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 282aa MW: 31013.3 Da PI: 8.7589 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 107.1 | 9.6e-34 | 24 | 78 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWtp+LHerFv+av++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
BGIOSGA017051-PA 24 KPRLRWTPDLHERFVDAVTRLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRL 78
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.502 | 21 | 81 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.4E-33 | 21 | 80 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.76E-16 | 24 | 80 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-24 | 24 | 79 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.9E-9 | 26 | 77 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.8E-21 | 125 | 169 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 282 aa Download sequence Send to blast |
MFEGCYGGVG SAAAAAAAAT RDPKPRLRWT PDLHERFVDA VTRLGGPDKA TPKSVLRLMG 60 MKGLTLYHLK SHLQKYRLGR QSKKSAGLEL AVADSGEFTA EGISFSIGAP PRNPAGGNNT 120 GEIPLADALK YQVEVQRKLQ EQLEVQKKLQ MRIEAQGRYL KEILEKAQKN ISLDANGSAN 180 LSSTRSQITD INLALSGFMD NATQVQEENN ELMKPISDDN LKVNNLGFQL YHLGSQESKD 240 VKCTPKTEEL LLLDLNIQGG YELSSRGMQG CELDLKINQQ RR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 2e-21 | 24 | 80 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-21 | 24 | 80 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-21 | 24 | 80 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-21 | 24 | 80 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.12937 | 0.0 | callus | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32991609 | 0.0 | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
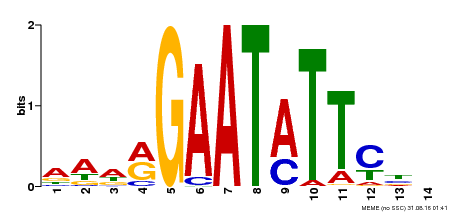
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK106400 | 0.0 | AK106400.1 Oryza sativa Japonica Group cDNA clone:002-102-G05, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015635409.1 | 0.0 | myb family transcription factor PHL11 isoform X2 | ||||
| TrEMBL | B8ATM6 | 0.0 | B8ATM6_ORYSI; Uncharacterized protein | ||||
| STRING | ONIVA04G23170.1 | 0.0 | (Oryza nivara) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3940 | 34 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 3e-53 | G2-like family protein | ||||



