 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_022020 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 425aa MW: 47120.6 Da PI: 9.3781 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 42.8 | 1.2e-13 | 346 | 398 | 5 | 57 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkeva 57
+r +r++kNRe+A rsR+RK+a++ eLe +v++L++ N++++++ ee ++ +
DCAR_022020 346 RRRKRMIKNRESAARSRARKQAYTLELEAEVAKLKEVNEEMQSKQEEFMEMQK 398
689*****************************************999998865 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 3.7E-11 | 342 | 406 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.277 | 344 | 395 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 4.5E-12 | 346 | 398 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 3.6E-12 | 346 | 395 | No hit | No description |
| CDD | cd14707 | 2.04E-23 | 346 | 400 | No hit | No description |
| SuperFamily | SSF57959 | 7.74E-10 | 346 | 395 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 349 | 364 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 425 aa Download sequence Send to blast |
MGSYINFKNY GVTSQTEGSG SKQAENPSLA RQSSVYSLTF EELQNTLDGV GKDFGSMNMD 60 ELLKSIWTAE GNQPVISSGD IRQGIPFPGG NLNKQGSLTL PRTLSQKTVD EVWKDLLKET 120 DNMRDGNGGM NLSEREATLS EMTLEEFLSR AGVVGGTTHF GRSYNGSHGD ITQQDNSNAD 180 LPFGFPQTEV SYESRSNNVI KNRNAVHNPY SSLGLNSGGI TPQQQPRQQQ QPLLPKQVTL 240 AFASPLNEGN NAQLTSLGTR SPLVRRGDPF MDNSATQSCV RQSREIGIAS LAPRVAIPVG 300 YHKSQLTSNM NPDRNLDASS LSHSPYAYNE GTNGRRTGTF EKVVERRRKR MIKNRESAAR 360 SRARKQAYTL ELEAEVAKLK EVNEEMQSKQ EEFMEMQKNQ ILEKMNMPWG SKRLCLRRTL 420 TGPW* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
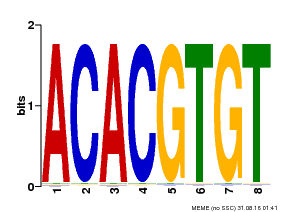
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017256174.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_017256176.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| Refseq | XP_017256177.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 7 | ||||
| TrEMBL | A0A164VNK2 | 0.0 | A0A164VNK2_DAUCS; Uncharacterized protein | ||||
| STRING | XP_009791697.1 | 1e-136 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA776 | 24 | 97 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G45249.1 | 3e-72 | abscisic acid responsive elements-binding factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_022020 |



