 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do007201.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 263aa MW: 29008 Da PI: 9.4693 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 105 | 6.2e-33 | 41 | 133 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
F+ k++++++d++++ +++w gn+f+vld+ f++ +Lp+yFkh+nfaSFvRQLn+YgF+kv++++ weF+h+sF +g+ +ll i
Do007201.1 41 FVAKTFHMVSDPATDAVVRWGGAGNTFLVLDPAAFSDYLLPSYFKHRNFASFVRQLNTYGFRKVDSDR---------WEFAHESFLRGQAHLLPLIV 128
9****************************************************************999.........******************** PP
XXXXX CS
HSF_DNA-bind 99 rkkse 103
rkk++
Do007201.1 129 RKKKK 133
**986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 4.0E-36 | 30 | 124 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 8.0E-50 | 37 | 130 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 5.03E-32 | 38 | 130 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 2.1E-17 | 41 | 64 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 8.6E-29 | 41 | 130 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.1E-17 | 79 | 91 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 80 | 104 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.1E-17 | 92 | 104 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006970 | Biological Process | response to osmotic stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MGSECKDQLQ LQHAGFGTGG RGGALQSQRQ QEEQAGAIAP FVAKTFHMVS DPATDAVVRW 60 GGAGNTFLVL DPAAFSDYLL PSYFKHRNFA SFVRQLNTYG FRKVDSDRWE FAHESFLRGQ 120 AHLLPLIVRK KKKSGGGREL FEEGEEVRGT IRDVRRLQEE QRGMEEELRA MDRRLRAAES 180 RPCQMMAFLA KLADDPRVVL RAMLAKEEEL AAGKGPLPVE APSKRRRIGV GAEAAGAGGE 240 AAELAQSRGA VPFPFSVLGQ VFY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ldu_A | 3e-25 | 38 | 130 | 17 | 120 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 223 | 227 | KRRRI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
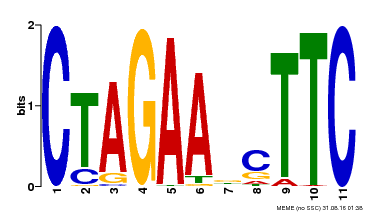
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do007201.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU954042 | 0.0 | EU954042.1 Zea mays clone 1452946 heat shock factor protein HSF30 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004969854.1 | 1e-134 | heat stress transcription factor C-1b | ||||
| Swissprot | Q942D6 | 1e-127 | HFC1B_ORYSJ; Heat stress transcription factor C-1b | ||||
| TrEMBL | A0A1E5W7Y1 | 0.0 | A0A1E5W7Y1_9POAL; Heat stress transcription factor C-1b | ||||
| STRING | Si002580m | 1e-134 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11298 | 31 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 1e-62 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



