 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Manes.08G005700.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Crotonoideae; Manihoteae; Manihot
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 513aa MW: 55917.8 Da PI: 9.6259 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 16.1 | 3.1e-05 | 65 | 87 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+ C++C+k F r nL+ H r H
Manes.08G005700.1.p 65 FICEICNKGFQRDQNLQLHRRGH 87
89*******************88 PP
| |||||||
| 2 | zf-C2H2 | 12.7 | 0.00038 | 142 | 164 | 1 | 23 |
EEETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCpdCgksFsrksnLkrHirtH 23
+kC++C+k + +s+ k H +t+
Manes.08G005700.1.p 142 WKCEKCSKKYAVQSDWKAHSKTC 164
58*****************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 64 | 87 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.36E-7 | 64 | 87 | No hit | No description |
| Pfam | PF12171 | 6.2E-5 | 65 | 87 | IPR022755 | Zinc finger, double-stranded RNA binding |
| SMART | SM00355 | 0.022 | 65 | 87 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.845 | 65 | 87 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 67 | 87 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 79 | 107 | 137 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 3.8E-5 | 130 | 163 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 5.36E-7 | 137 | 162 | No hit | No description |
| SMART | SM00355 | 140 | 142 | 162 | IPR015880 | Zinc finger, C2H2-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0045604 | Biological Process | regulation of epidermal cell differentiation | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 513 aa Download sequence Send to blast |
MQMMSGDAFS IPSSIPLFAP EEQNANPNPK TNPVPKKKRN LPGTPDPDAE VIALSPKTLM 60 ATNRFICEIC NKGFQRDQNL QLHRRGHNLP WKLRQRTNKE VIRKKVYICP EKTCVHHDPS 120 RALGDLTGIK KHYSRKHGEK KWKCEKCSKK YAVQSDWKAH SKTCGTREYK CDCGTLFSRK 180 DSFITHRAFC DALAEESARI SSVPATNLNF RNDTVNFPRG EPAGGGVQDI AGISQFSSGF 240 RPDFNGFPSL SADQQKTGLS LWLNQANSQI NPADIVPNHA NLYASSSSTG LPEMVQMGSN 300 LYGSSSTTNF GNLKLSGLPH GLKEEEGSNK GLNMVDSLPS LYSDSHQNRQ SKPAAPMSAT 360 ALLQKAAQMG STRSNQSFFG SNSHGLMSSS SSSNTTNLNT LSQNRNELHQ VFQNVNKQPE 420 CNITVTSSVA MGDAIMVVSS GLDQVAMQSS GRQNDPFQLK VQPGSTSLES GLTRDFLGMS 480 AQSGRPFLPQ ELAKFASMSS AMGLSQFTNN PN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5b3h_C | 4e-38 | 138 | 207 | 3 | 72 | Zinc finger protein JACKDAW |
| 5b3h_F | 4e-38 | 138 | 207 | 3 | 72 | Zinc finger protein JACKDAW |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 91 | 104 | KLRQRTNKEVIRKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
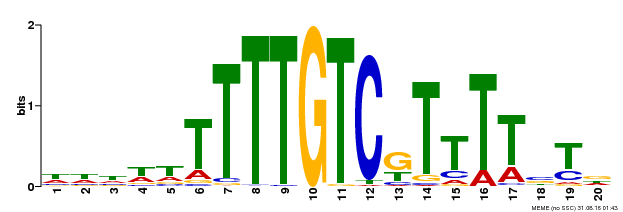
| MP00485 | DAP | Transfer from AT5G03150 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Manes.08G005700.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681841 | 6e-64 | LN681841.1 Cucumis melo genomic scaffold, anchoredscaffold00011. | |||
| GenBank | LN713258 | 6e-64 | LN713258.1 Cucumis melo genomic chromosome, chr_4. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021619709.1 | 0.0 | zinc finger protein JACKDAW-like | ||||
| TrEMBL | A0A2C9VC91 | 0.0 | A0A2C9VC91_MANES; Uncharacterized protein | ||||
| STRING | cassava4.1_032876m | 0.0 | (Manihot esculenta) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF181 | 34 | 275 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G03150.1 | 1e-107 | C2H2-like zinc finger protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Manes.08G005700.1.p |



