 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Mapoly0043s0098.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Marchantiophyta; Marchantiopsida; Marchantiidae; Marchantiales; Marchantiaceae; Marchantia
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 740aa MW: 77773.8 Da PI: 7.226 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 72.9 | 4e-23 | 160 | 261 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f k+lt+sd+++ g +++p+ +ae++ +++++ +t+ +d +g++W++++iyr++++r++lt+GW+ Fv++++L +gD +vF
Mapoly0043s0098.1.p 160 FAKTLTQSDANNGGGFSVPRYCAETIfppldySIDPPV-QTVLAKDVHGERWKFRHIYRGTPRRHLLTTGWSTFVNQKKLVAGDAIVFL- 247
89****************************99555554.49************************************************. PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
++ ++el+v+v+r+
Mapoly0043s0098.1.p 248 -RTASGELCVGVRRS 261
.458999*****985 PP
| |||||||
| 2 | Auxin_resp | 102.6 | 4.9e-34 | 361 | 443 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
aa+ a ++++FevvY+Prast+eF+vk++ v++al++ + +GmRfkmafetedss++++ +Gt+++v+ +d+ Wp+S+Wr+L+
Mapoly0043s0098.1.p 361 AASLAVQGQPFEVVYYPRASTAEFCVKAQAVKAALDHTWFPGMRFKMAFETEDSSRISWfMGTISAVQPADS-LWPKSPWRVLQ 443
6778899**************************************************************999.9********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 2.4E-36 | 152 | 271 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 3.66E-30 | 156 | 272 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.46E-20 | 159 | 260 | No hit | No description |
| SMART | SM01019 | 2.9E-19 | 160 | 262 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 6.9E-21 | 160 | 261 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.675 | 160 | 262 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.0E-29 | 361 | 443 | IPR010525 | Auxin response factor |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 740 aa Download sequence Send to blast |
MPGPSPGCGT MSGTNIKMEK SEESMGGGGK GWGGGRDRDS SSDGGGGSGE NTGLDPQLWH 60 ACAGGMVQLP PVGAKVIYFP QGHGEQAATP PEFPRMMGPQ GTIGCRVVSV SFLADTETDE 120 VYARIRLQPL EREAAMSIAD STLDADGGPS SPPPEKPASF AKTLTQSDAN NGGGFSVPRY 180 CAETIFPPLD YSIDPPVQTV LAKDVHGERW KFRHIYRGTP RRHLLTTGWS TFVNQKKLVA 240 GDAIVFLRTA SGELCVGVRR SMRGTGGADS STWSGGSSTS HHRPNRWEVK GTESFSDFLG 300 NDSAAGGGSV SSAGSAAGPG GPRAGPGGSN SGPGIGIPGP STTSSFARNR ARVTAQSVLE 360 AASLAVQGQP FEVVYYPRAS TAEFCVKAQA VKAALDHTWF PGMRFKMAFE TEDSSRISWF 420 MGTISAVQPA DSLWPKSPWR VLQVTWDEPD LLQGVSRVSP WQVELVSTLP MQLPPFSLPK 480 KKLRAAQPSD MNMQGQGLMG MPITLPSVFG QINPWAHGLT MEEVTAGMQG ARHDRVFGLA 540 LSEKFRPGKL PGGFFSAAEG YYPDHAGRGG GGEPHLSAFP LQDRASNISS LISSLGSVPP 600 SGDHGSAGAL LVPAGSWSGG SANNKSASTQ LVLFGQAINT DSNKSQPQHS GGSSCDGPSL 660 QHLKEESSGK RKDSPSESNQ NENFERGQKY MSGSLSNGKQ SDLIVGEFQK WVPGLDKERG 720 AGEKLAASPE ILQSPQGAW* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldu_A | 2e-56 | 54 | 464 | 51 | 388 | Auxin response factor 5 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 27 | 34 | GGKGWGGG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
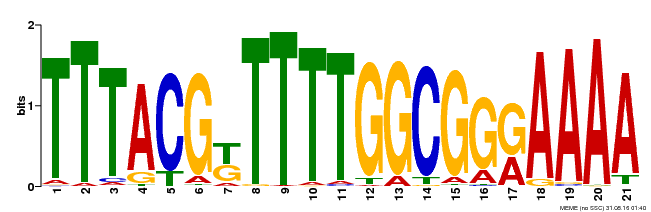
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP877971 | 0.0 | KP877971.1 Marchantia polymorpha auxin response factor 3 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024372940.1 | 0.0 | auxin response factor 10-like isoform X1 | ||||
| Refseq | XP_024372941.1 | 0.0 | auxin response factor 10-like isoform X1 | ||||
| Swissprot | Q653H7 | 1e-166 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A0G3FJH3 | 0.0 | A0A0G3FJH3_MARPO; Auxin response factor | ||||
| STRING | PP1S339_47V6.1 | 0.0 | (Physcomitrella patens) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP526 | 15 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G28350.1 | 2e-92 | auxin response factor 10 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Mapoly0043s0098.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



