 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | OGLUM01G45850.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 694aa MW: 73921.4 Da PI: 9.31 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 28.4 | 2.9e-09 | 408 | 455 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR++i +++ L++l+P + +k + Ka +L + ++Y++sLq
OGLUM01G45850.1 408 SHSLAERVRREKISQRMKLLQDLVPGC----NKVVGKAVMLDEIINYVQSLQ 455
79************************9....677*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00083 | 5.99E-10 | 402 | 459 | No hit | No description |
| SuperFamily | SSF47459 | 2.36E-16 | 402 | 473 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.256 | 404 | 454 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 2.0E-16 | 406 | 472 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 2.3E-6 | 408 | 455 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-10 | 410 | 460 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 694 aa Download sequence Send to blast |
MTEILSPTKT ILLADCSSKK AHAFNPRKGG GCKDLQQQQQ QLVLLFHSFT VPPLFLALSR 60 TRKAPAFTLS PLAFPHPRRS GIWWLLASRV SFAGLGRSWE LGNSGGLVGF RGELRQIFEM 120 NCGPPDQLPP ATAPSCFLNL NWDQSMDAAA GGHLDPALSS MVSSPASNST GALHGISPQP 180 HYGGGTPLSS PPKLNLSMMG QFHHYAAPPQ VGGGGGGGGG GGLPILENLM PMGHLDQFLA 240 DPGFAERAAR LSGFDARGGG GGGGYGGAGP AQFGLPDAGA AGASKEMELG NTRDESSVSD 300 PAPGGAEIPP KGASDGNARK RKASGKGKGK DSPMSTSAAK EDSSGKRCKS TEESNAAAEE 360 NSGKGKAAQS NSENGGGKKQ GKDSSSKPPE PPKDYIHVRA RRGEATDSHS LAERVRREKI 420 SQRMKLLQDL VPGCNKVVGK AVMLDEIINY VQSLQRQVEF LSMKLATVNP QLDFNNLPNL 480 LAKDMHQSCS PLQSSHFPLE TSGAPLPYIN QPQQGNPLGC GLTNGMDNQG SMHPLDPAFC 540 RPMGSHHPFL NGVSDAASQV GAFWQDDLQS VVQMDMGQGQ EIATSSNSYN VLEGSENKLN 600 FRICLSQKRV HMKIQDYLTA SPADFFSTIS MGIDNVMRYK KETALQQMRR SHGKKRGAKG 660 ETGTVHIHSH KQSIIKPCLR LLADNFYSRR DIQS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00660 | PBM | Transfer from Pp3c6_9520 | Download |
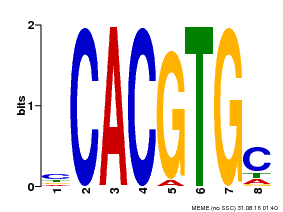
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK101063 | 0.0 | AK101063.1 Oryza sativa Japonica Group cDNA clone:J033022O19, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015621926.1 | 0.0 | transcription factor bHLH77 isoform X1 | ||||
| TrEMBL | A0A0D9YJ49 | 0.0 | A0A0D9YJ49_9ORYZ; Uncharacterized protein | ||||
| STRING | OGLUM01G45850.1 | 0.0 | (Oryza glumipatula) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP919 | 38 | 137 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G07340.1 | 3e-57 | bHLH family protein | ||||



