 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pahal.G00294.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Panicinae; Panicum
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 661aa MW: 73110.2 Da PI: 5.6707 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 77 | 2e-24 | 128 | 228 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrse 89
f+k+lt sd++++g +++ +++aee+ ++++++ ++l+ +d++g +W +++i+r++++r++lt+GW+ Fv++++L +gD ++F+ + ++
Pahal.G00294.1 128 FCKTLTASDTSTHGGFSVLRRHAEEClpqldmSQNPPC-QELVAKDLHGTEWHFRHIFRGQPKRHLLTTGWSVFVSSKRLVAGDAFIFM--RGEN 219
99*****************************7556666.59************************************************..4499 PP
E..EEEEE- CS
B3 90 felvvkvfr 98
+el+v+v+r
Pahal.G00294.1 220 GELRVGVRR 228
99*****97 PP
| |||||||
| 2 | Auxin_resp | 99.6 | 4.2e-33 | 254 | 339 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsd...ldpvrWpnSkWrsLk 83
a+ha+st++ F+v+Y+Pr+s+s+F+v+v+k+++a k+k+svGmRfkm+fe+++++err+sGt++++ ++ + W +S WrsLk
Pahal.G00294.1 254 ASHAISTGTLFSVFYKPRTSRSDFIVSVNKYLEAKKQKISVGMRFKMRFEGDEAPERRFSGTIIDIGSlpaMSKSLWADSDWRSLK 339
789*************************************************************997622267778********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 2.75E-44 | 115 | 257 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 1.4E-41 | 125 | 243 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 9.26E-22 | 126 | 228 | No hit | No description |
| PROSITE profile | PS50863 | 11.716 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 5.7E-22 | 128 | 228 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.6E-21 | 128 | 230 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 2.1E-31 | 254 | 339 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.9E-12 | 542 | 638 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 27.002 | 549 | 642 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 5.23E-12 | 551 | 628 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0042802 | Molecular Function | identical protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 661 aa Download sequence Send to blast |
MAAAMDVANP GAAAGSGMAS DALYRELWHA CAGPLVTVPR QGERVYYFPQ GHMEQLEAST 60 HQQLDQYLPM FNLPSKILCS VVNVELRAEA DSDEVYAQIM LQPEADQSEL TSPDPELKEP 120 EKCTAHSFCK TLTASDTSTH GGFSVLRRHA EECLPQLDMS QNPPCQELVA KDLHGTEWHF 180 RHIFRGQPKR HLLTTGWSVF VSSKRLVAGD AFIFMRGENG ELRVGVRRLM RQVNSMPSSV 240 ISSHSMHLGV LATASHAIST GTLFSVFYKP RTSRSDFIVS VNKYLEAKKQ KISVGMRFKM 300 RFEGDEAPER RFSGTIIDIG SLPAMSKSLW ADSDWRSLKV QWDEPSSILR PDRISPWEVE 360 PLDAANPQSP QPPLRNKRAR PPASPSTVAE LPSGFGLWKS PTDSARTLSF SDPQRARELF 420 PSIPTSTFSS SSNVNFNSKN EPSMLNSQFY WSARDTRADS CAASTNAVIV EKKQEQNSGG 480 CRLFGIDICS AEEEVLPVVT APALGYDQTA ASLELNSDKL SQPSDVNNSD APGASSERSP 540 PESQSRQVRS CTKVIMQGMA VGRAVDLTKL SGYSDLCHKL EEMFDIQGEL GSTLKKWRVI 600 YTDDEDDMML VGDDPWNEFC DMVKRIYIYT YEEAKKLTSK SNLPGSSDTS KLSDVNSQSE 660 * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 0.0 | 5 | 364 | 1 | 357 | Auxin response factor 1 |
| 4ldw_A | 0.0 | 5 | 364 | 1 | 357 | Auxin response factor 1 |
| 4ldw_B | 0.0 | 5 | 364 | 1 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
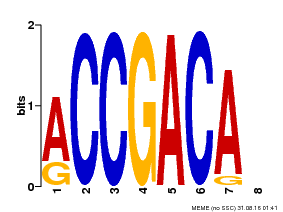
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00097 | PBM | Transfer from AT1G59750 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025823850.1 | 0.0 | auxin response factor 9-like | ||||
| Swissprot | Q0JCZ4 | 0.0 | ARFI_ORYSJ; Auxin response factor 9 | ||||
| TrEMBL | A0A2S3I6V4 | 0.0 | A0A2S3I6V4_9POAL; Auxin response factor | ||||
| STRING | Pavir.Gb01617.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1082 | 36 | 116 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G59750.2 | 0.0 | auxin response factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Pahal.G00294.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



