 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Phvul.002G238400.1 | ||||||||
| Common Name | PHAVU_002G238400g | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Phaseolus
|
||||||||
| Family | Dof | ||||||||
| Protein Properties | Length: 184aa MW: 20989.5 Da PI: 7.9356 | ||||||||
| Description | Dof family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-Dof | 123.1 | 9.7e-39 | 43 | 100 | 3 | 60 |
zf-Dof 3 ekalkcprCdstntkfCyynnyslsqPryfCkaCrryWtkGGalrnvPvGggrrknkk 60
e++++cprC+s++tkfCy+nny+++qPr+fCk+C+ryWt+GGalrnvP+G+grrk k
Phvul.002G238400.1 43 ERIIPCPRCKSMETKFCYFNNYNVNQPRHFCKGCQRYWTAGGALRNVPIGAGRRKAKP 100
67899**************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| ProDom | PD007478 | 8.0E-29 | 41 | 99 | IPR003851 | Zinc finger, Dof-type |
| Pfam | PF02701 | 1.1E-31 | 45 | 99 | IPR003851 | Zinc finger, Dof-type |
| PROSITE profile | PS50884 | 28.155 | 46 | 100 | IPR003851 | Zinc finger, Dof-type |
| PROSITE pattern | PS01361 | 0 | 48 | 84 | IPR003851 | Zinc finger, Dof-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010214 | Biological Process | seed coat development | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 184 aa Download sequence Send to blast |
MAQVESGQAS EGIKLFGTTI TLHGRERRED NKVSEDETEK RAERIIPCPR CKSMETKFCY 60 FNNYNVNQPR HFCKGCQRYW TAGGALRNVP IGAGRRKAKP PSHDPTAFLD ASTVQQQFAL 120 EEWDVATITH GDFRHFFPSK RRRINSAEHE SQNEFVMSHE IIEHPNEIET KEALKGSDNV 180 KLI* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 139 | 143 | KRRRI |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds specifically to a 5'-AA[AG]G-3' consensus core sequence. Acts as a negative regulator in the phytochrome-mediated light responses. Controls phyB-mediated end-of-day response and the phyA-mediated anthocyanin accumulation. Not involved in direct flowering time regulation. {ECO:0000269|PubMed:19619493}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00166 | DAP | Transfer from AT1G29160 | Download |
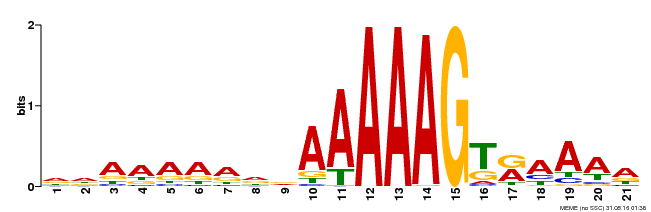
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Phvul.002G238400.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By red or far-red light. Circadian-regulation. The transcript level rises progressively from dawn and decreases during the night. {ECO:0000269|PubMed:19619493}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP015034 | 1e-126 | AP015034.1 Vigna angularis var. angularis DNA, chromosome 1, almost complete sequence, cultivar: Shumari. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007159448.1 | 1e-137 | hypothetical protein PHAVU_002G238400g | ||||
| Swissprot | P68350 | 4e-53 | DOF15_ARATH; Dof zinc finger protein DOF1.5 | ||||
| TrEMBL | V7CML4 | 1e-136 | V7CML4_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007159448.1 | 1e-136 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF6483 | 32 | 51 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G29160.1 | 1e-55 | Dof family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Phvul.002G238400.1 |
| Entrez Gene | 18638926 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



