 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Spipo4G0037700 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Araceae; Lemnoideae; Spirodela
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 542aa MW: 60877.6 Da PI: 5.8879 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 81.3 | 1.1e-25 | 196 | 249 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r++W+ +LH++Fv+av+q+ G +k Pk+il+lm+v+gLt+e+v+SHLQkYRl
Spipo4G0037700 196 KARVVWSVDLHQKFVNAVNQI-GFDKVGPKKILDLMDVPGLTRENVASHLQKYRL 249
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 82.3 | 1.6e-27 | 17 | 125 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaG 93
vl+vdD+p+ +++l+++l+k y ev++++ ++ al+ll+e++ +D+++ D++mp+mdG++ll++ e +lp+i+++ ge + + + ++ G
Spipo4G0037700 17 VLVVDDDPTWLKILEKMLRKCSY-EVTTCSLARVALDLLRERKdwFDIVISDVNMPDMDGFKLLEQVGLEM-DLPVIMMSVDGETSRVMKGIQHG 109
89*********************.***************888888**********************6644.8********************** PP
ESEEEESS--HHHHHH CS
Response_reg 94 akdflsKpfdpeelvk 109
a d+l Kp+ ++el++
Spipo4G0037700 110 ACDYLLKPVRMKELRN 125
*************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.40.50.2300 | 1.3E-44 | 14 | 151 | No hit | No description |
| SuperFamily | SSF52172 | 8.97E-36 | 14 | 146 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 1.4E-32 | 15 | 127 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.755 | 16 | 131 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 1.5E-24 | 17 | 126 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 1.81E-30 | 18 | 131 | No hit | No description |
| SuperFamily | SSF46689 | 7.79E-18 | 193 | 253 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.8E-28 | 195 | 254 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-22 | 196 | 249 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.0E-7 | 198 | 248 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 542 aa Download sequence Send to blast |
MENSSSRDES FPAGLRVLVV DDDPTWLKIL EKMLRKCSYE VTTCSLARVA LDLLRERKDW 60 FDIVISDVNM PDMDGFKLLE QVGLEMDLPV IMMSVDGETS RVMKGIQHGA CDYLLKPVRM 120 KELRNIWQHV FRKKIHEVRE TENYDSYEDN NIRSRLDELD ENLVFNGAFH SSKKRKDNHY 180 REHMEQSFGD SVAVKKARVV WSVDLHQKFV NAVNQIGFDK VGPKKILDLM DVPGLTRENV 240 ASHLQKYRLY LSRLQKQNEE KTLGGAAQTD LNAKEPPGVL SQQSSLARQP KSSPSGYGFS 300 STRIYTQDMI PKIAAFRATE TELASNDEAR VSPKKGNITR FGLMASKNLE PWGDGGNQLT 360 QFVQFMKGQQ SVSVKDLLQF RQAGQDHRFE HDHAQSPHIV NLRGSIEAEG TSPSVQMKPA 420 DSKEGKYMLL KCGLDSGSSQ ISHGFVNAHS FQPASITNLS CGNPCSSDDK NSIGSWSSLY 480 SDAEAPPSID SLTFPLHGLG AFECNQIASD FQSILYDELQ YNCEYLSDFF EAPMVDDGLF 540 T* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 5e-17 | 193 | 254 | 2 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
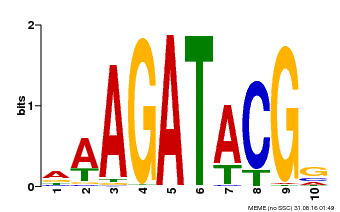
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010932292.1 | 1e-163 | two-component response regulator ORR26 | ||||
| Swissprot | Q5N6V8 | 1e-150 | ORR26_ORYSJ; Two-component response regulator ORR26 | ||||
| TrEMBL | A0A2H3YDP8 | 1e-156 | A0A2H3YDP8_PHODC; Two-component response regulator | ||||
| STRING | XP_008797265.1 | 1e-156 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5900 | 37 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-115 | response regulator 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Spipo4G0037700 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



