 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_016647439.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Amygdaleae; Prunus
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 431aa MW: 48305.1 Da PI: 6.38 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 85 | 6e-27 | 158 | 222 | 1 | 70 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwk 70
r+++sL+llt+kf++l++++++++++ln+ a+ L +v ++RRiYDi+NVLe++nliek++kn+irwk
XP_016647439.1 158 RYDSSLGLLTKKFVSLIQEAKDRTLDLNKTAEVL---EV--QKRRIYDITNVLEGINLIEKTSKNHIRWK 222
6899******************************...99..****************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 1.8E-27 | 157 | 222 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 4.6E-33 | 158 | 223 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| SuperFamily | SSF46785 | 4.22E-17 | 158 | 221 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF02319 | 1.7E-24 | 160 | 222 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 4.67E-33 | 234 | 342 | No hit | No description |
| SuperFamily | SSF144074 | 5.49E-23 | 235 | 341 | No hit | No description |
| Pfam | PF16421 | 4.0E-23 | 240 | 339 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000902 | Biological Process | cell morphogenesis | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042023 | Biological Process | DNA endoreduplication | ||||
| GO:0051782 | Biological Process | negative regulation of cell division | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046982 | Molecular Function | protein heterodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 431 aa Download sequence Send to blast |
MEMEDPNRLL RPSQFEFQLL HSHSQTQNHQ NHPSYLLPSL LSSQHLLPHS FPAFAPSIDS 60 GHHRLSDSAF ASHADAHSAF AKLALRPKKE IDDVAQIGGH EAAAGPSKVL HDPSLSIPQS 120 CTGRKHNPKS KVPKHTKAGT QRSTADSHNG LNPAIGCRYD SSLGLLTKKF VSLIQEAKDR 180 TLDLNKTAEV LEVQKRRIYD ITNVLEGINL IEKTSKNHIR WKCYEGSTPG ELDDQVSRIQ 240 DEVESLYAEE CRLDDAIREK LELLRALEED EKNKKFLFLT EEDILSLCCF QNQTLIAIKA 300 PQASYIEVID PDDEDTCFPQ KQFRMIVRST IGPIHLYLLS KYQGQRENIA VKQAKSVGSS 360 SWNSSDYSRV EDGGLSSYHQ GNKKDSSEIL SLPGSEASSG IQKIIPSHVN ANDDYWFRSE 420 PEVSLTDLWA N |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 9e-23 | 153 | 222 | 2 | 72 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00190 | DAP | Transfer from AT1G47870 | Download |
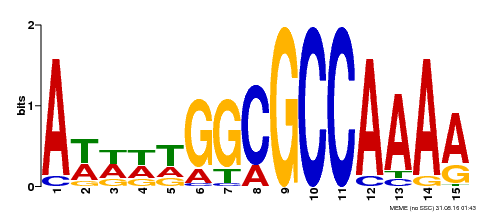
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | XP_016647439.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016647439.1 | 0.0 | PREDICTED: transcription factor E2FC | ||||
| TrEMBL | A0A314ZVN9 | 0.0 | A0A314ZVN9_PRUYE; Transcription factor E2FC | ||||
| STRING | XP_008221406.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF11885 | 25 | 29 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G47870.2 | 1e-86 | E2F/DP family protein | ||||



