 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma304g00070.1 | ||||||||
| Common Name | ZOSMA_304G00070 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 259aa MW: 28993.4 Da PI: 4.6006 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 62.1 | 1.3e-19 | 86 | 135 | 3 | 56 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkklege 56
y+G+r+++ +g+W+AeIrdp++ r++lg+f taeeAa+a++ +k+++ge
Zosma304g00070.1 86 YRGIRQRP-WGKWAAEIRDPRK---GVRVWLGTFNTAEEAARAYDSEAKRIRGE 135
9*******.**********954...4**************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.7E-22 | 85 | 143 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 9.02E-34 | 85 | 142 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 4.2E-32 | 85 | 143 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.775 | 85 | 142 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.1E-37 | 85 | 148 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.0E-12 | 86 | 97 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 5.0E-13 | 86 | 135 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 6.0E-12 | 108 | 124 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008219 | Biological Process | cell death | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010286 | Biological Process | heat acclimation | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051707 | Biological Process | response to other organism | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 259 aa Download sequence Send to blast |
MCGGAIISGF IPPSESLTAI LADDPLDADF DADFEADFED FNDDSGEDES SAVKPLAFAT 60 RSEISATVTV LKYGNTGRAR KRKNIYRGIR QRPWGKWAAE IRDPRKGVRV WLGTFNTAEE 120 AARAYDSEAK RIRGEKAKLN FPKTLPYAVS KLNRKRKSLD NMMTCNSQHN KKTEKLQKNS 180 DEVVASAVPI FFDETETVNA VKLLSDEDEI SSYEHPYMMM IMPCPEESSS ELGSNDLQVY 240 EDSLWNFDDT PIFGNLAY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1gcc_A | 1e-22 | 86 | 142 | 3 | 60 | ETHYLENE RESPONSIVE ELEMENT BINDING FACTOR 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250, ECO:0000269|PubMed:9159183}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
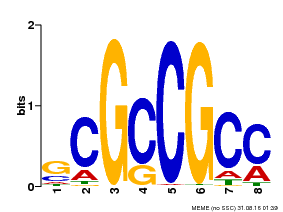
| Motif ID | Method | Source | Motif file |
| MP00109 | PBM | Transfer from AT3G16770 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By 1-aminocyclopropane-1-carboxylic acid (ACC, ethylene precursor), methyl jasmonate (MeJA), and Botrytis cinerea. Also induced by cadmium. {ECO:0000269|PubMed:18836139, ECO:0000269|PubMed:9159183}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_007139068.1 | 3e-51 | hypothetical protein PHAVU_009G262200g | ||||
| Swissprot | P42736 | 1e-39 | RAP23_ARATH; Ethylene-responsive transcription factor RAP2-3 | ||||
| TrEMBL | A0A0K9PA37 | 0.0 | A0A0K9PA37_ZOSMR; Uncharacterized protein | ||||
| STRING | XP_007139068.1 | 1e-50 | (Phaseolus vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16770.1 | 1e-37 | ethylene-responsive element binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma304g00070.1 |



