 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_6581_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 105aa MW: 11947.8 Da PI: 8.9888 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 58.3 | 1.7e-18 | 31 | 78 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
cra_locus_6581_iso_2_len_410_ver_3 31 KGPWTPEEDIILVSYIQEHGPGNWRSVPTNTGLLRCSKSCRLRWTNYL 78
79******************************99************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 7.2E-25 | 22 | 81 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 25.075 | 26 | 82 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.4E-14 | 30 | 80 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.8E-17 | 31 | 78 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 3.22E-22 | 32 | 105 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 3.23E-12 | 33 | 78 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-7 | 82 | 105 | IPR009057 | Homeodomain-like |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010118 | Biological Process | stomatal movement | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 105 aa Download sequence Send to blast |
KSTHKVEEAE EKEIIINMGR PPCCDKVGIK KGPWTPEEDI ILVSYIQEHG PGNWRSVPTN 60 TGLLRCSKSC RLRWTNYLRP GIKRGNFTPH EEGMIIHLQA LLGNK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the regulation of gene (e.g. drought-regulated and flavonoid biosynthetic genes) expression and stomatal movements leading to negative regulation of responses to drought and responses to other physiological stimuli (e.g. light). {ECO:0000269|PubMed:22018045}. | |||||
| UniProt | Transcription factor involved in the regulation of gene (e.g. drought-regulated and flavonoid biosynthetic genes) expression and stomatal movements leading to negative regulation of responses to drought and responses to other physiological stimuli (e.g. light) (PubMed:16005291, PubMed:21637967). Promotes guard cell deflation in response to water deficit. Triggers root growth upon osmotic stress (e.g. mannitol containing medium) (PubMed:21637967). {ECO:0000269|PubMed:16005291, ECO:0000269|PubMed:21637967}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00132 | DAP | Transfer from AT1G08810 | Download |
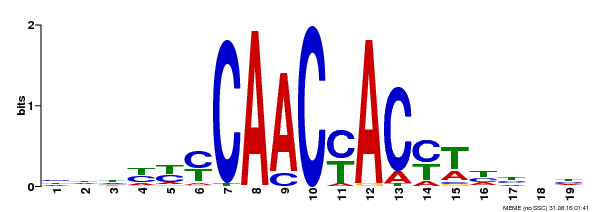
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by jasmonic acid (JA) and salicylic acid (SA) (PubMed:16463103). Triggered by auxin (IAA) in roots (PubMed:9839469, PubMed:21637967). Stimulated by light, UV-light and cold (PubMed:9839469). According to PubMed:16005291 and PubMed:23828545, rapidly repressed by abscisic acid (ABA) in an ABI1-dependent manner. But in contrast, according to PubMed:21637967, transiently induced by ABA in seedlings (PubMed:16005291, PubMed:21637967, PubMed:23828545). Rapidly repressed by drought. Activated by white light, but repressed by blue light and darkness (PubMed:16005291). Transiently induced by salt (NaCl) in seedlings. Induced by sucrose (PubMed:21637967). Positively regulated by SCAP1 (PubMed:23453954). {ECO:0000269|PubMed:16005291, ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:21637967, ECO:0000269|PubMed:23453954, ECO:0000269|PubMed:23828545, ECO:0000269|PubMed:9839469}. | |||||
| UniProt | INDUCTION: Repressed by abscisic acid (ABA) and osmotic stress (salt stress). {ECO:0000269|PubMed:22018045}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006484863.1 | 2e-62 | myb-related protein 306 | ||||
| Refseq | XP_012465520.1 | 2e-62 | PREDICTED: myb-related protein 306-like | ||||
| Swissprot | B3VTV7 | 3e-62 | MYB60_VITVI; Transcription factor MYB60 | ||||
| Swissprot | Q8GYP5 | 3e-62 | MYB60_ARATH; Transcription factor MYB60 | ||||
| TrEMBL | A0A0D2VHT3 | 4e-61 | A0A0D2VHT3_GOSRA; Uncharacterized protein | ||||
| STRING | Gorai.013G196800.1 | 7e-62 | (Gossypium raimondii) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



